ITK 5.3 Release Candidate 1 available for testing
We are happy to announce the Insight Toolkit (ITK) 5.3 Release Candidate 1 is available for testing! ITK is an open-source, cross-platform toolkit for N-dimensional scientific image processing, segmentation, and registration.
ITK 5.3 is a feature release that accelerates performance, provides new segmentation and shape analysis algorithms, and makes over 200 more improvements.
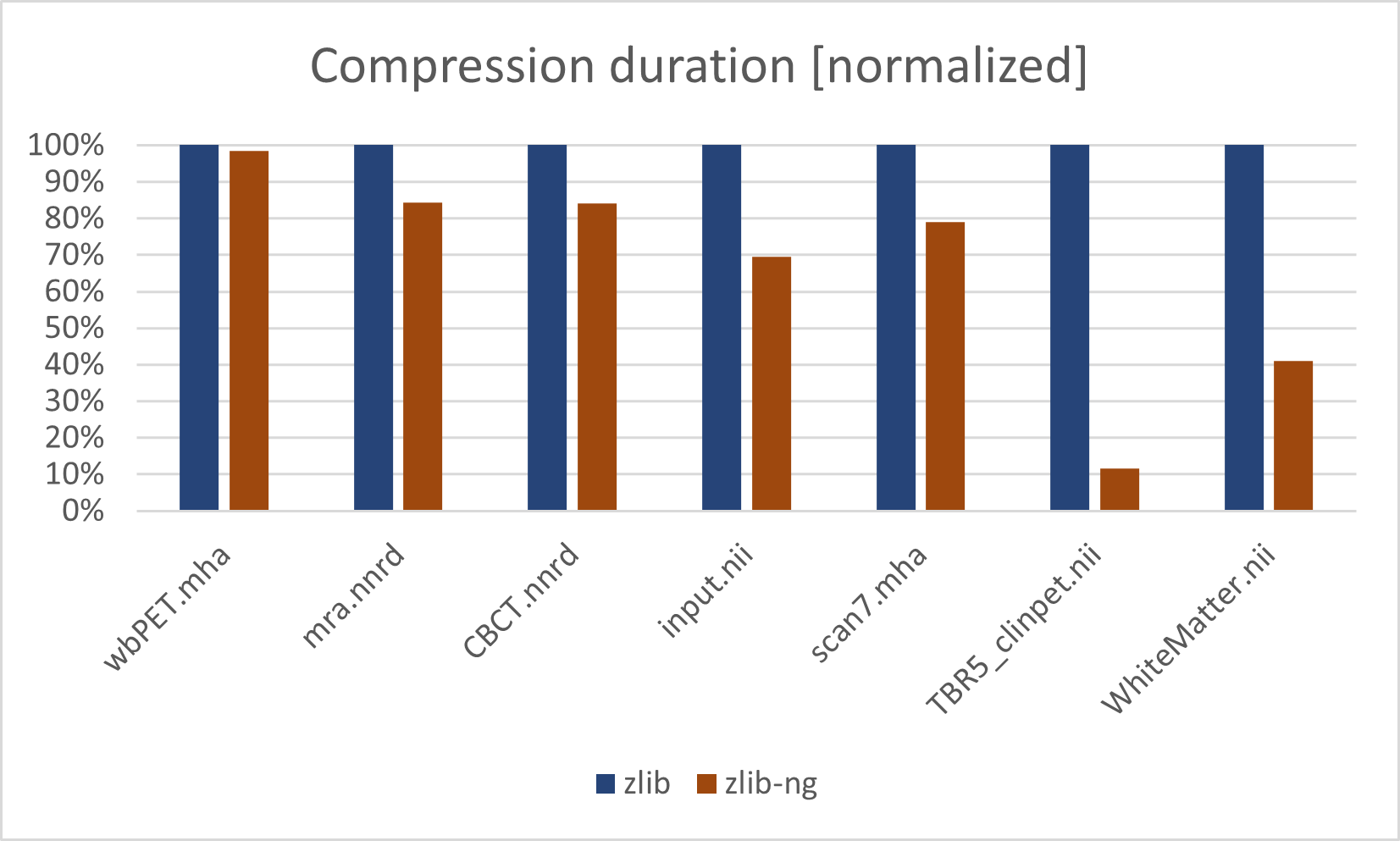
Release Candidate 1 highlights performance improvements. For deformable image registration, b-spline sampling was improved by ~30%. Multi-threading in Python was improved by adding Threading Building Blocks (oneTBB) to the cross-platform binaries, which improves multi-threaded parallelism by ~5-10%. Compression time with zlib, used by common medical imaging file formats like NIFTI, NRRD, or MetaImage, was dramatically reduced through migration to zlib-ng.

| name | description | zlib duration [ms] | zlib C.Ratio | zlib-ng duration [ms] | zlib-ng C.Ratio | zlib-ng Speed-up | zlib-ng C. Ratio improvement |
|---|---|---|---|---|---|---|---|
| wbPET.mha | whole body PET | 672 | 9.7% | 661 | 9.3% | 2% | 4% |
| mra.nrrd | MR angiography | 1300 | 49.0% | 1097 | 49.1% | 19% | 0% |
| CBCT.nrrd | ConeBeam CT | 11281 | 42.5% | 9486 | 41.3% | 19% | 3% |
| input.nii | brain MRI | 4818 | 57.6% | 3353 | 57.4% | 44% | 0% |
| scan7.mha | mouse ultrasound | 5939 | 45.4% | 4694 | 46.2% | 27% | -2% |
| TBR5_clinpet.nii | label map | 782 | 0.5% | 91 | 0.5% | 759% | -2% |
| WhiteMatter.nii | label map | 185 | 5.6% | 76 | 5.8% | 143% | -4% |
| Average | 145% | 0% |
Comparison of image compression with traditional zlib library and the new zlib-ng replacement introduced with ITK 5.3 using the default compression level.
Download
Python Packages
Install ITK Python packages with:
pip install --upgrade --pre itkLibrary Sources
Testing Data
Unpack optional testing data in the same directory where the Library Source is unpacked.
Checksums
Features
Python
- Python packages now include oneTBB support for improved performance.
- Following CPython’s deprecation schedule Python 3.6 is no longer supported.
- Initial Python wrapping is available for the Video modules.
TransformToDisplacementFieldis now available in Python.
C++
- C++14 is now required.
- The minimum CMake version required is now 3.16.3.
- New functions:
MakePoint,MakeVector,MakeIndex,MakeSize.
New filter
itk::TransformGeometryImageFilter: applies a rigid transform to anImage‘s metadata.
Remote module updates
New remote modules:
- HASI: High-Throughput Applications for Skeletal Imaging
- ITKGrowCut: segments a 3D image from user-provided foreground and background seeds
Updated modules: AdaptiveDenoising, AnisotropicDiffusionLBR, BSplineGradient, BoneEnhancement, BoneMorphometry, Cuberille, GrowCut, HASI, HigherOrderAccurateGradient, IOFDF, IOScanco, IsotropicWavelets, MinimalPathExtraction, Montage, MorphologicalContourInterpolation, RTK, SimpleITKFilters, SkullStrip, SplitComponents, Strain, TextureFeatures, Thickness3D, TotalVariation, TubeTK, and Ultrasound.
Third party library updates
- gdcm
- niftilib
- zlib migrated to zlib-ng
- hdf5
- kwsys
- metaio
- googletest
- vxl
Congratulations
Congratulations and thank you to everyone who contributed to this release.
Of the 32 authors who contributed since v5.2.0, we would like to specially recognize the new contributors:
Pranjal Sahu, Darren Thompson, Tomoyuki SADAKANE, Oleksandr Zavalistyi, Jose Tascon, Kian Weimer, Michael Kuczynski, Ebrahim Ebrahim, and Philip Cook.
What’s Next
We anticipate an additional release candidate following community testing before the 5.3.0 release. The following release candidates will improve related documentation and make further improvements. Please try out the current release candidate, and discuss your experiences at discourse.itk.org. Contribute with pull requests, code reviews, and issue discussions in our GitHub Organization.
Enjoy ITK!