Medical Researchers Can Integrate, Visualize, and Analyze Metabolomics Data Using Viime, Kitware’s All-Inclusive, Multi-Platform Solution
Viime: A multi-platform solution
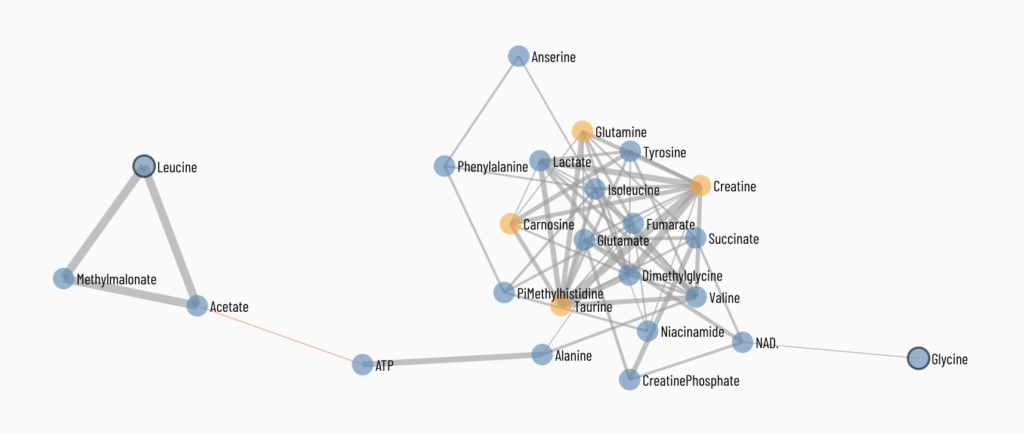
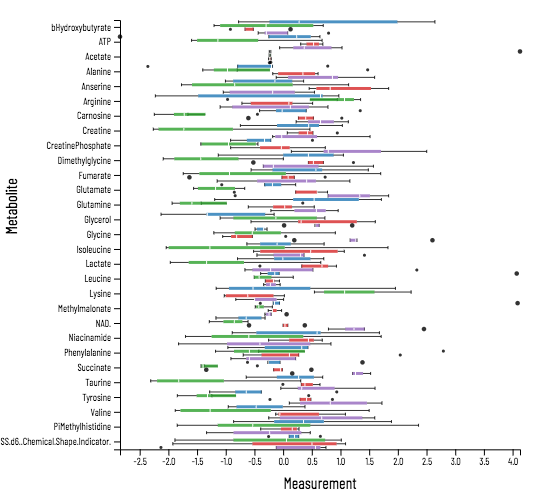
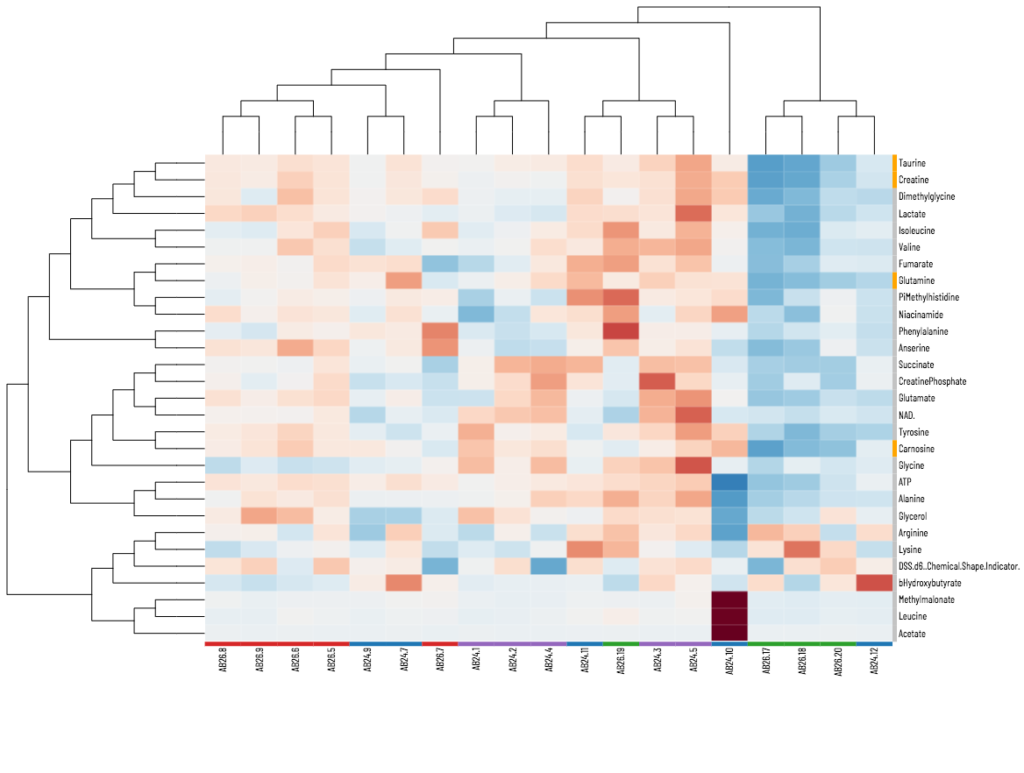
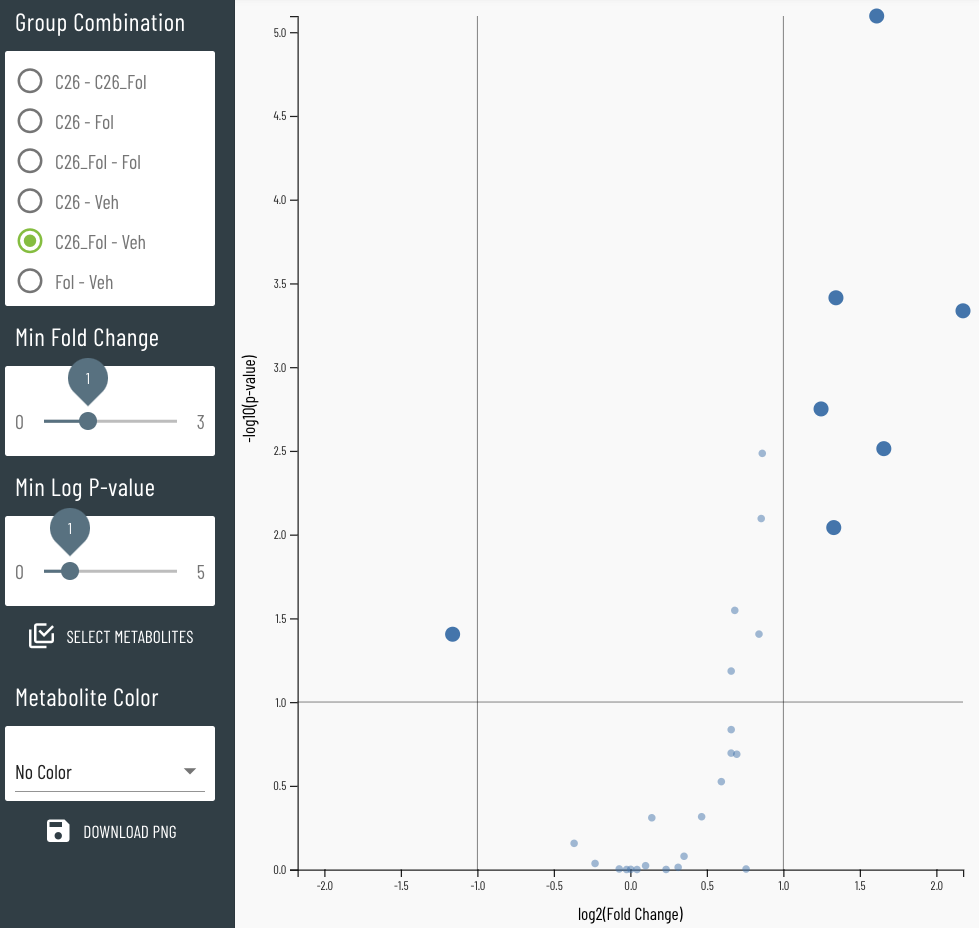
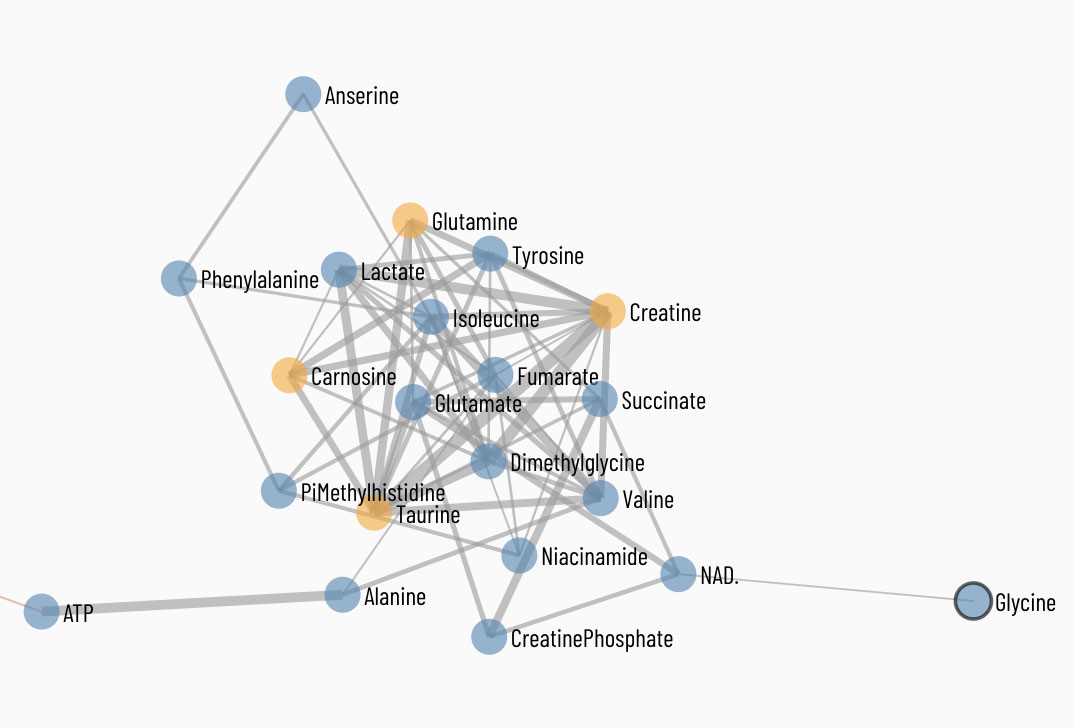
Viime (VIsualization and Integration of Metabolomics Experiments) is an open source web application that provides a new workflow for metabolomics research by offering state-of-the-art integration algorithms and visualizations. Viime pulls together metabolomics data from different sources, integrates the data, and enables analysis and visualization. These interactive tables, charts, heatmaps, and network diagrams allow medical researchers to gain a thorough understanding of their experimental data (see Figs. 1-4).
Metabolomics involves the comprehensive measurement of metabolites from a biological system. Factors such as genetics, lifestyle, biological stressors, disease, diet, and the environment can influence these metabolomic profiles. Medical researchers have to rely on multiple platforms to collect data that is all-inclusive. This multi-platform approach yields diverse data structures, so integrating the data can be challenging. Instead of analyzing datasets separately, which prevents the observation of potentially interesting relationships and correlations between metabolites, Viime easily integrates metabolomics data from multiple platforms.
The Viime Workflow
Medical scientists who use Viime have the flexibility to upload their data files in a semi-automated process (see Fig. 5) and get immediate visual feedback. The feedback is interactive, so the results of various ingestion, pretreatment, and analysis options are represented in real-time (see Fig. 6). Scientists can also run algorithms to obtain more accurate results using data from measurement platforms or tissues, highlight metabolites that should be further investigated, and export data and charts.
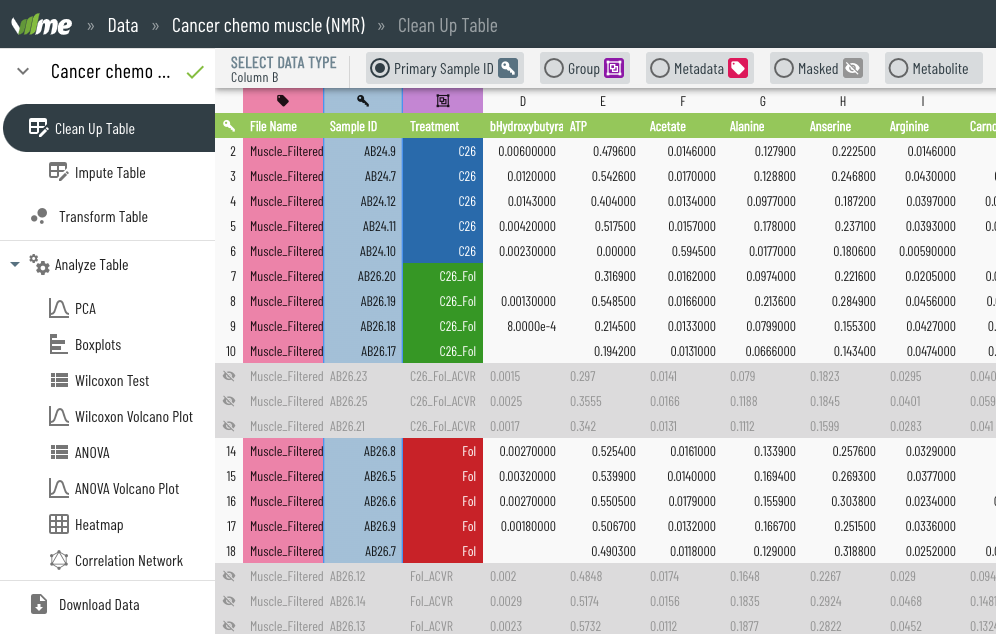
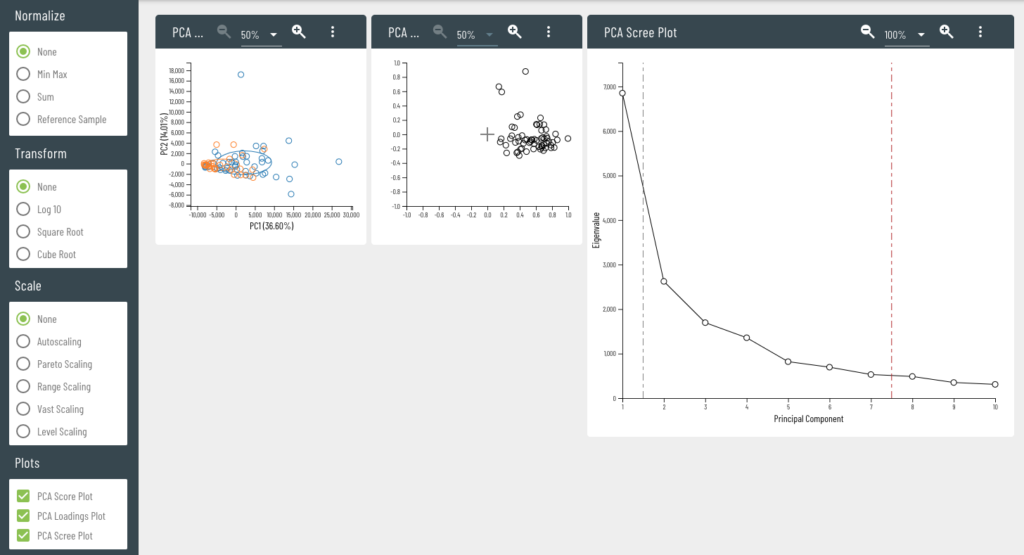
These features help medical scientists understand the role various metabolites may play in certain medical conditions. This research is essential for evaluating drug therapies and their effect on patients—for example, cachexia, which appears in some cancer patients but not in others.
Viime in the Medical Research Industry
Kitware’s Data and Analytics Team developed Viime as part of the National Institute of Health’s (NIH) Metabolomics Data Integration Phase 2 project. Kitware received assistance from Thomas O’Connell, Lilian Golzarri Arroyo, and Jun Wan from Indiana University, and Daniel Raftery from the University of Washington. Kitware’s work on Viime has been published in the Journal of Open Source Software (JOSS) under the paper titled Viime: Visualization and Integration of Metabolomics Experiments.
Leveraging Viime to Advance Your Research
If you are interested in using Viime for your medical research, contact us at kitware@kitware.com. The software is open source and can be used without restriction, but partnering with Kitware will ensure that you are using the application effectively and getting the results you need. We will work closely with you to develop a custom solution tailored to your specific requirements. Let us help you unlock the full potential of your data.
Our Data and Analytics Team has created a platform of tools to enhance the way customers use data. These tools have proven valuable across multiple disciplines from scientific research, to engineering design, to software development. With scalable, web-based data management, powerful, intuitive analytics engines, and functionality to create graphs, charts, geospatial maps, and other visualizations, these tools are designed to enable powerful custom solutions for any data need. We partner with universities on government-funded projects and with commercial companies on product development. Learn more about the capabilities of our Data and Analytics Team by visiting Kitware’s Data and Analytics page.
This project has been funded in whole or in part with Federal funds from the National Cancer Institute, National Institutes of Health, Department of Health and Human Services, under Contract No. HHSN261201800044C (UPIID 75N91018C00044). The percentage of the total costs of the project that was financed with Federal money is 100%. The dollar amount of Federal funds for the project is $1,443,759.