VTK HDF Reader
Motivation
HDF is a widely used data format which consists of n-dimensional named arrays (datasets) organized in a directory structure (groups). Storing VTK data in HDF is complicated because HDF does not have a standard way to attach meaning to its arrays – so a reader does not know which array represents points, cells, and other attributes. Traditionally, storing VTK data in HDF has been done using XDMF. However, this method also has some significant drawbacks:
- The user needs to write an additional XML file describing the mapping between the HDF arrays and the VTK data structure.
- XDMF is not actively maintained.
- There are two versions of XDMF, version two and version three, with version three not fully implementing version two.
A fully supported option is to use ADIOS2, which can save data as HDF and use Fides for describing the VTK data model. While this option offers in-situ and in-transit functionality and the option to switch to a different binary format (BP5 binary-pack), it introduces complexity that may not be desired for a developer looking for a simple solution to save their data in HDF.
To address these issues, we have designed a VTK HDF Data Format and implemented a reader for this format. Our main objective is to make it easy to write, from a parallel simulation, data that can be processed in VTK. Currently we support serial and parallel image data and unstructured grid and serial Overlapping AMR support. For a usage example and sample datasets see the unit testing for the reader. Note that we use only one format for serial or parallel processing and only one extension for all VTK data formats. Additional VTK data types would be added depending on interest and funding. This reader is available in VTK and ParaView master.
Overview
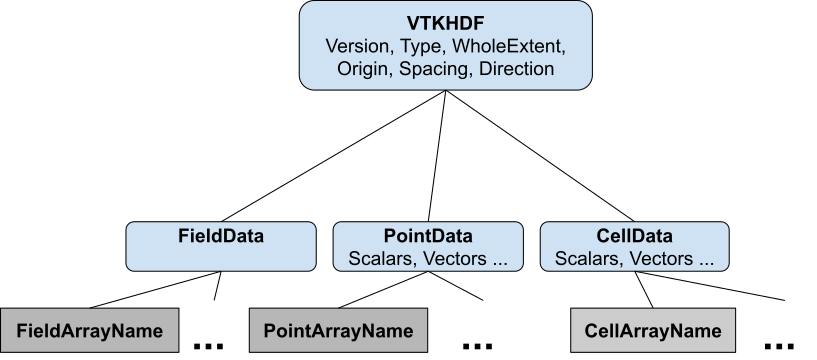
The VTK HDF file format for the image data set is shown in Figure 1. In this figure, groups are shown with rounded blue rectangles. Inside the rectangle, we have the group name as a title and the group attributes underneath the tile. Datasets are shown with gray rectangles.

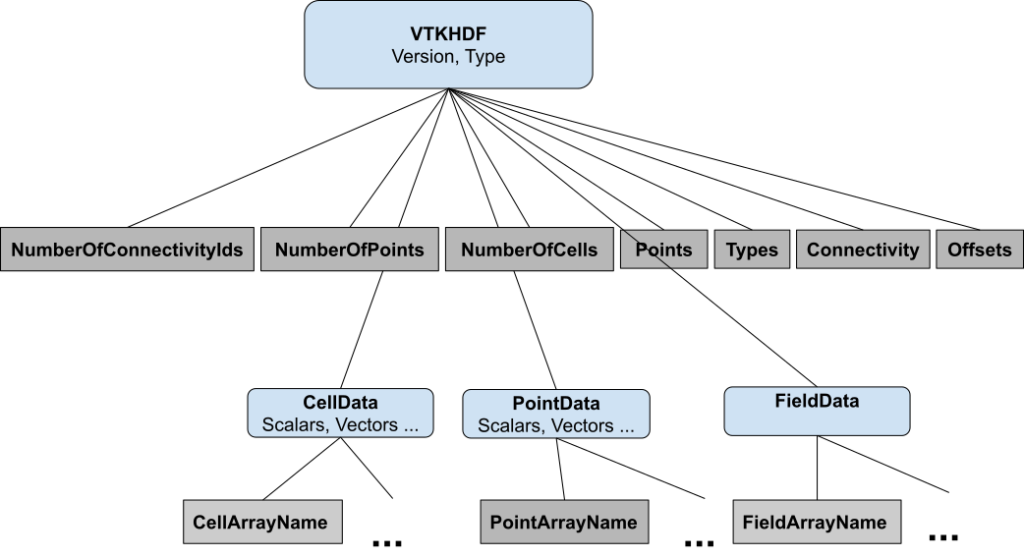
Attribute arrays for the image dataset are not split into pieces for parallel processing. For ease of processing, we use the HDF hyper-slab API instead and rely on chunking to improve performance. The VTK HDF file format for an unstructured grid is shown in Figure 2. For parallel processing, attribute, point, and cell arrays are split into pieces, which are being recorded using NumberOfConnectivityIds, NumberOfPoints, and Number of Cells datasets. For instance, NumberOfPoints is a dataset that stores in entry i, the number of points for piece i.
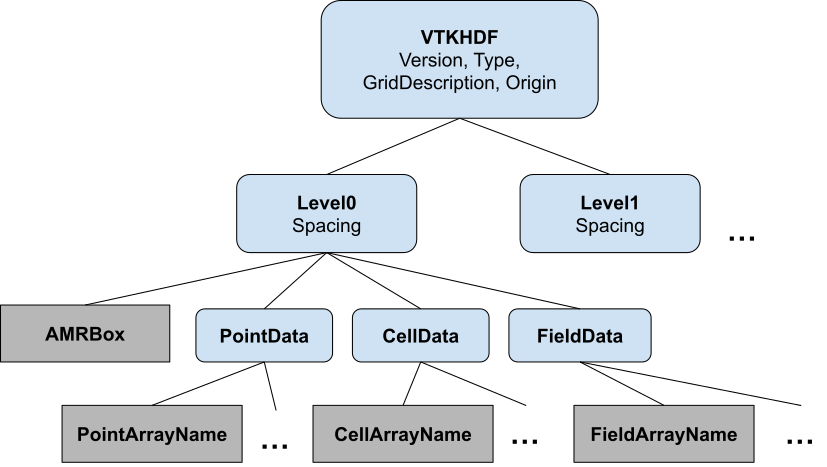
In designing this specification and reader we used a VTK XML to VTK HDF converter written in Python, which may prove useful to developers wishing to save their data as VTK HDF. The VTK HDF file format for Overlapping AMR is detailed in Figure 3. Each level in an overlapping AMR file format (and data structure) consists of a list of uniform grids with the same spacing (as specified by the Spacing attribute). The AMRBox dataset contains the bounding box for each of these grids. The points and cell arrays for these grids are stored serialized in one dimension and stored in a dataset in the PointData or CellData group. For a detailed description of these formats see VTK File Formats on the VTK Examples website.
Conclusion
With this specification and the reader, we provide a simple and direct way to save data in the HDF format that can be natively read into VTK. We look forward to hearing feedback on this work on the VTK Discourse website.
Acknowledgements
The work presented in this article was funded by ORNL ECP Alpine and KM Turbulenz GmbH, km-turbulenz.de
Very nice, do you want to do an example for VTK Examples website?
Maybe using can-pvtu.hdf? You can also link to https://gitlab.kitware.com/danlipsa/vtkxml-to-vtkhdfin the documentation. I would suggest it goes in vtk-examples/src/Python/IO/
Let me know if I can help.
Hi Andrew, Sure I’ll look into that.
Do you intend to provide a writer for this new format?
It would be great if the format could describe temporal dataset. Is it something you have in mind for later or not?
KEU is looking into implementing a writer. We deal with temporal data in the same way with other standard VTK datatypes through vtkFileSeriesReader. That should already work in ParaView.
is there a place where I could download an working example HDF5 file in this format?
Building VTK with VTK_BUILD_TESTING=WANT will create tests for the HDF format. You can run these tests using ctest -R HDFReader. Searching in TestHDFReader.cxx for .hdf will show you a number of VTKHDF files created in the build directory by the testing framework.
For your information, the VTKHDF file format roadmap has been published in discourse : https://discourse.vtk.org/t/vtkhdf-roadmap/13257
The VTKHDF Roadmap has a new location in the VTK documentation: https://docs.vtk.org/en/latest/vtk_file_formats/vtkhdf_file_format/vtkhdf_roadmap_status.html