Tailored Applications are Major Trend at 3D Slicer 2016 Winter Project Week

 Earlier this month, we attended the 3D Slicer 2016 Winter Project Week at the Massachusetts Institute of Technology (MIT) Computer Science and Artificial Intelligence Laboratory (CSAIL) in Cambridge, MA. The project week is a semi-annual event where 3D Slicer users and developers gather to discuss and work collaboratively on open-science solutions for problems that occur at the intersection of the computer science, mechanical engineering, biomedical engineering, and medical fields.
Earlier this month, we attended the 3D Slicer 2016 Winter Project Week at the Massachusetts Institute of Technology (MIT) Computer Science and Artificial Intelligence Laboratory (CSAIL) in Cambridge, MA. The project week is a semi-annual event where 3D Slicer users and developers gather to discuss and work collaboratively on open-science solutions for problems that occur at the intersection of the computer science, mechanical engineering, biomedical engineering, and medical fields.
During the week, we joined approximately 80 attendees—mainly from the U.S., with some traveling from Canada, Germany, Norway, and France—to work on close to 50 projects covering image-guided therapy (IGT), image analysis, tractogaphy, and the 3D Slicer infrastructure. We made progress on specific issues; helped out the 3D Slicer community; and discussed design, a roadmap, and key priorities.

Overall, the consensus at the project week was to maintain a lean, robust, and well-documented 3D Slicer core that can be easily extended. This quote by Ron Kikinis from Brigham and Women’s Hospital (BWH) illustrates this point: “A well-maintained core is needed for currently funded applications to fully leverage 3D Slicer as a platform and to protect NIH’s investment.”
Additional highlights are as follows:
-
We discussed the Tracked Ultrasound Standardization effort. The goal of this effort is to establish a common software platform for tracked ultrasound and image-guided interventions by converging existing IGT toolkits including 3D Slicer and the SlicerIGT extension, the Plus toolkit, CustusX, Intraoperative Brain Imaging System (IBIS), Echelon, and Medical Imaging Interaction Toolkit-IGT (MITK-IGT). Lead by our friends at Queen’s University, a draft implementation of the effort is expected by June 2016.
-
For SlicerOpenCV, Nicole Aucoin from BWH is working on a 3D Slicer extension using an ITK external module to build and package the ITKOpenCV module into its own extension. ITKOpenCV is a core ITK module that is neither built nor packaged in the 3D Slicer core. Instead, it is built externally within its own extension.
-
Docker is gaining a lot of traction. Many are trying to make sense of it within the context of medical imaging research. Coupling it with solutions based on HTML5 virtual network computing (VNC) clients like noVNC has potential. It is worth noting two things:
-
Folks in Australia are deploying an infrastructure at the national level that leverages Docker, named DIT4C.
-
Kitware team member Matt McCormick created SlicerDocker and docker-opengl projects.
-
-
This year, the project week tutorial contest winner was Csaba Pinter from Queen’s University, who explained how to use the Subject Hierarchy module. The purpose of the tutorial contest is “to enrich the training materials that are available to end-users and developers using 3D Slicer.”
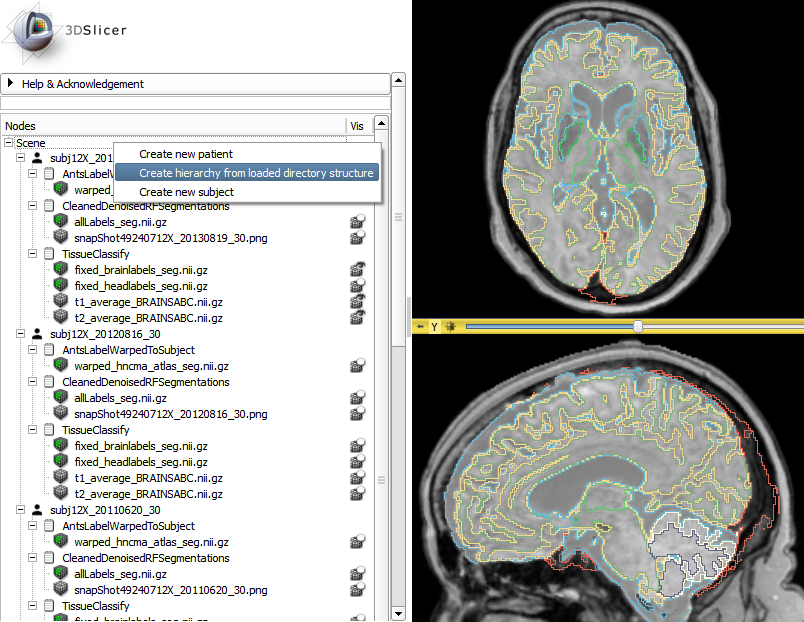
In other findings, the trend across the 3D Slicer community is to develop tailored applications geared toward specific tasks. There are a few approaches that can be used to achieve this:
-
At Kitware, we have an internal process for efficiently creating tailored applications. The recent post “Introducing NIRFAST-Slicer 1.0” provides a great example.
-
Otherwise, for concept validation and quick prototyping, the Slicelets approach could be considered. It consists of 3D Slicer started without its main window and a Python script to create a simple user interface (UI). The lumpectomy (breast conserving tumor excision) Slicelet from our collaborators at Queen’s University is a good example. Note that the video in the link showcases 3D Slicer running on a 32-bit Surface tablet.
-
The 3D Slicer community has worked on another approach, named Custom Slicer Generator. Steve Pieper is the main architect of that project. François Budin, who joined Kitware recently, used and improved it to package a special version of 3D Slicer (SlicerCMF, from the Dental and Craniofacial Bionetwork for Image Analysis). It consists of helper scripts that allow you to create archives containing the 3D Slicer core along with a selection of 3D Slicer modules and extensions.
What is great is that these approaches are all complementary, as they contribute to building reusable blocks.
To wrap up this post, we will leave you with some food for thought. 3D Slicer is no longer a monolithic infrastructure; it is a modular framework that allows you to either create professional-scale applications or quickly prototype ideas. Similar to what ParaView is for scientific computing, 3D Slicer is your tool for medical imaging. Both ParaView and 3D Slicer allow you to share your creative and efficient solutions built on top of CMake, the Common Toolkit (CTK), the Insight Segmentation and Registration Toolkit (ITK), Python, or the Visualization Toolkit (VTK).
If you would like help with adapting 3D Slicer to your projects, please contact Jean-Christophe Fillion-Robin via kitware@kitware.com.