Beyond Affine: Thin Plate Splines for Serial Histology Alignment
When Your Images Don’t Quite Line Up
If you work with serial histology slides, you are familiar with the routine: Zoom into a perivascular region. Toggle to the adjacent stain. Pan. Nudge. Recenter. Repeat.
Serial sections are routinely used for cross-stain interrogation and volumetric reconstruction. But consecutive sections from the same tissue block rarely align well enough for direct comparison, especially at high magnification. Sections may be skipped, and tissue deforms during cutting and mounting in ways that undermine direct spatial correspondence.
Those distortions may seem small, but repeated hundreds of times a day, they compound. Misalignment undermines comparative analysis, annotation quality, and downstream machine learning workflows. At first glance, this seems like a straightforward fitting problem that a simple affine transform should be able to handle. But, once applied, this is clearly insufficient.
Why Simple Alignment Fails
A global affine transform can provide coarse alignment, but it assumes linear deformation across the entire image. In practice, deformation is anything but linear. One region may align well, while another may stretch, compress, or drift. These are serial sections, not the same section that has just been re-imaged. The farther you move from the region driving the fit, the less reliable the alignment becomes.
For detailed visual comparison or quantitative analysis, that inconsistency undermines both confidence and efficiency. What’s needed is a transformation that adapts locally while remaining globally smooth.
A Better Model for Image Adjustment
A more suitable model for this type of deformation is the thin plate spline (TPS) transformation. By anchoring corresponding landmarks between sections, TPS interpolation enforces exact alignment at known control points while smoothly adjusting surrounding tissue. The deformation adapts to local variation without introducing discontinuities or unrealistic warping.
This approach is well-established in image registration workflows and supported by common scientific imaging libraries.
The Scaling Problem
A 40x whole slide image can reach 10 gigapixels. At 0.25 micrometers per pixel, you’re navigating enormous spatial detail.
Alignment is rarely a one-pass operation. You place fiducials, evaluate, adjust, add more landmarks, and refine. Rewriting gigapixel images at every iteration introduces latency at precisely the moment you need rapid feedback.
Dynamic Registration in Practice
The HistomicsTK platform makes this process straightforward and interactive. Fiducial landmarks can be placed directly in the standard viewer interface. These point annotations define correspondence between sections.
Instead of copying and rewriting gigapixel images, a lightweight z-stack can be generated that references the original images along with their warp parameters. The viewer automatically exposes a slider control to move across sections. Tiles are fetched from the source images and warped dynamically on demand—no precalculation needed.
If only a few fiducial points are specified, the system gracefully falls back to simpler transforms. Registration quality can be evaluated immediately, without generating new full-resolution image files.
The outcome is a stable anatomical context across stains. You can examine a perivascular region in one slide, move to the next stain, and remain spatially grounded. That stability supports both interactive exploration and downstream computational analysis, freeing you to focus on the science.
Kitware’s Approach
At Kitware, we focus on solving exactly these kinds of problems, building reproducible registration workflows, implementing adaptive transformation models, and enabling interactive alignment at scale. We’ve done it in digital pathology, in geospatial systems, and in places where spatial integrity is critical. Details like alignment should disappear into the background; if your attention can’t remain on the biology, we should talk.
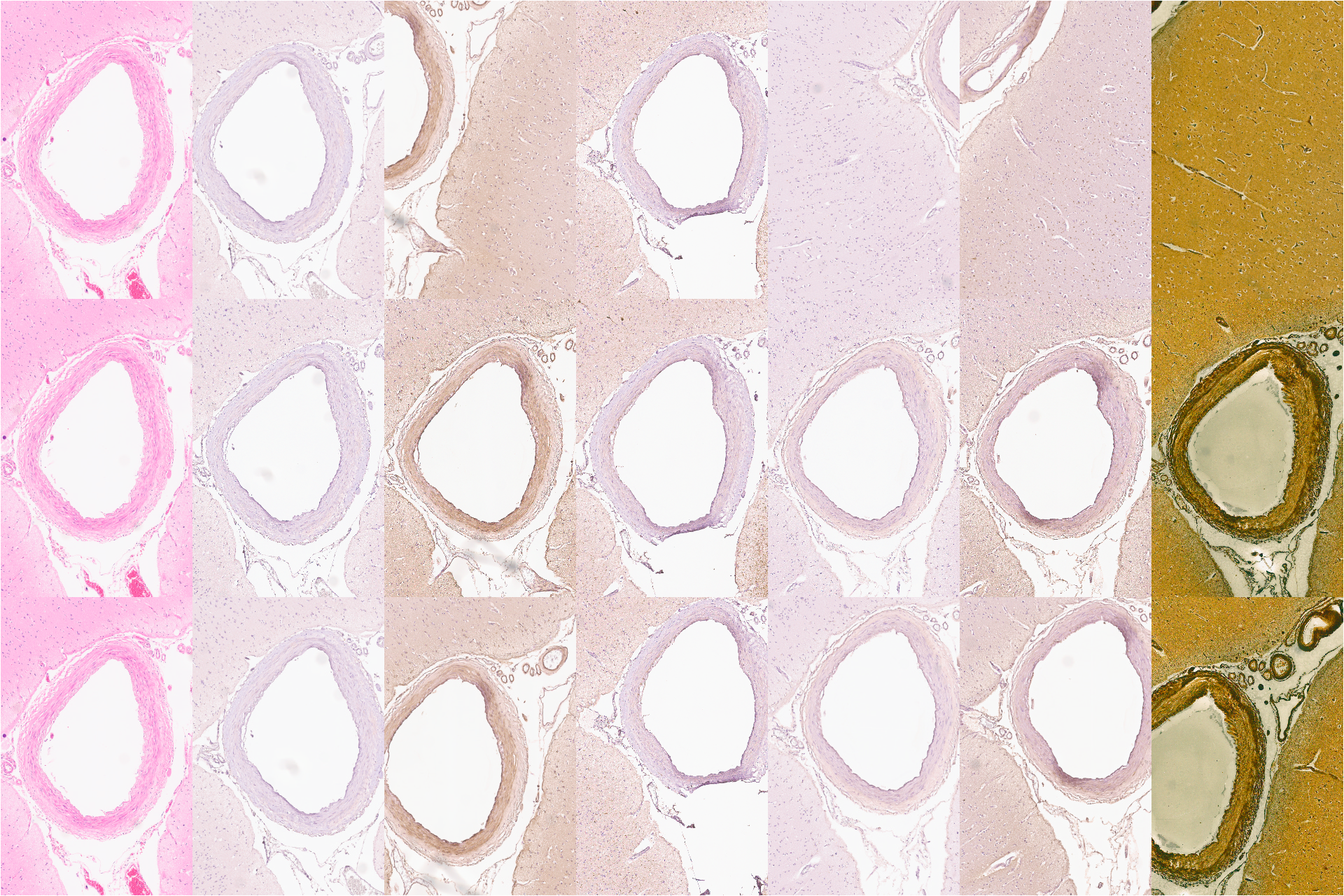
This video shows how alignment gives context. The left most column is the same for each row and shows an H&E stained image of some brain tissue. The top row is a set of serial sections as produced by the scanner (aside from de-identification). The bottom row are the same images aligned using an affine transform. The center row uses TPS (thin-plate spline) alignment, illustrating how context is preserved between sections.
Build Better Image Workflows With Kitware
Aligning serial histology sections becomes difficult when tissue deformation, skipped slices, and gigapixel images prevent reliable comparison. Kitware builds scalable registration workflows using adaptive transformation models such as thin plate splines and interactive landmark-based alignment. If your team is working with complex image data and needs reliable image registration at scale, contact Kitware to learn how we can help.