Designing an Open-source Python Framework for Large-Scale Diffusion MRI Analysis
Motivation: A unified framework for dMRI analysis
Diffusion MRI (dMRI) is a powerful non-invasive technique that uses the diffusion of water molecules to reveal the microstructural complexity of biological tissues. By measuring how cellular structures, such as membranes and myelin, constrain water movement, dMRI probes brain anatomy at a micrometer scale. This has made it an indispensable tool for mapping white matter integrity and connectivity, providing important insights into neurodevelopmental disorders, traumatic brain injury, and neurodegenerative diseases.
To extract quantitative insights from raw dMRI datasets, a typical analysis workflow involves several steps, including preprocessing, model fitting (such as DTI or microstructural models), and possibly tractography. A wide variety of specialized software libraries and tools have been developed to handle these specific tasks. However, for researchers or clinicians, this fragmented landscape, ranging from FSL for preprocessing to MRtrix3, DIPY, or AMICO for fiber modeling and microstructural indices, can create a significant barrier to analysis. This challenge is often compounded by the complex environment setups and installations required for each package.
To address this, we have developed a Python-native, pip-installable library, kwneuro, designed to unify commonly used methods into a cohesive ecosystem. By providing a standardized interface for various dMRI algorithms, this toolkit aims to simplify the transition from raw data to clinical insight, allowing researchers to explore datasets and perform complex analyses more easily. While not intended to replace powerful existing toolboxes, this library offers a lightweight alternative for researchers who need quick access to standard dMRI analysis methods.
Challenges in the Current dMRI Tooling Landscape
We identified several practical considerations in the current dMRI software landscape that can create barriers for exploratory and lightweight analyses:
- Installation and deployment complexity: Tools like FSL and MRtrix3 require binary installations with system-level dependencies, which can create challenges in diverse computing environments, from local workstations to institutional HPC clusters.
- Cross-Tool Integration: Transitioning between software environments (e.g., from FSL’s C++/shell-based tools to Python/MATLAB-based packages like AMICO) involves managing incompatible dependencies, paths, and file formats.
- Workflow Overhead: While Nipype provides robust workflow orchestration capabilities across multiple neuroimaging packages and facilitates reproducible pipeline definitions, users still face the burden of installing and configuring each underlying software package and must learn each tool’s distinct syntax and conventions.
- Input/Output Inconsistency: Inconsistent naming and storage conventions for inputs and outputs (scalar maps, tract labels, templates) across tools necessitate manual data management.
Design Principles: Accessibility and Ease of Use
kwneuro builds upon existing diffusion imaging tools, offering a user-friendly Python interface optimized for lightweight, exploratory analyses:
- Simple installation: Install via pip and start analyzing data in minutes, no complex dependencies or system configuration required.
- Flexible workflows: Our framework requires minimal configuration and makes it easy to test out different reconstruction models (e.g., from DTI to NODDI) without rewriting your entire analysis script.
- Standardized Artifacts: By enforcing consistent assumptions about inputs and outputs, our library ensures that your results are organized and ready for cohort-level statistical analysis.
- Transparent Data Handling: Control easily and precisely when data is loaded into memory or kept on disk for efficient processing.
Installation
pip install kwneuroExample workflow
from kwneuro.dwi import Dwi
from kwneuro.io import FslBvalResource, FslBvecResource, NiftiVolumeResource
dwi = Dwi(
NiftiVolumeResource(data_dir / f"{basename}.nii.gz"),
FslBvalResource(data_dir / f"{basename}.bval"),
FslBvecResource(data_dir / f"{basename}.bvec"),
)
dwi_denoised = dwi.denoise()
mask = dwi_denoised.extract_brain()
dti = dwi_denoised.estimate_dti(mask=mask)
fa_vol, md_vol = dti.get_fa_md()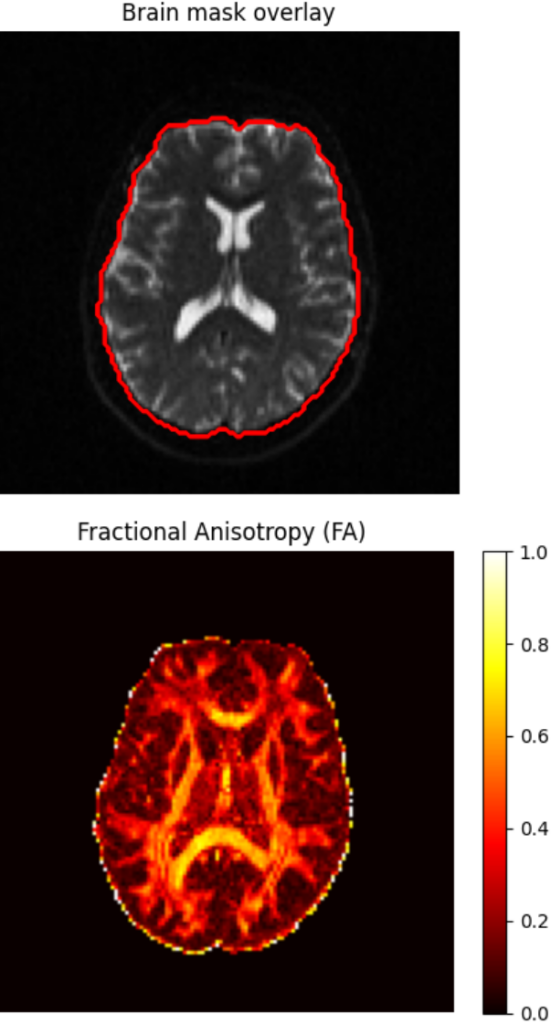
Example workflow demonstrating library installation and diffusion tensor fitting to compute a Fractional Anisotropy (FA) map. The streamlined interface requires minimal configuration to produce standard diffusion metrics.
Key Capabilities
The library provides a high-level Python interface for commonly used dMRI analysis tasks. Below is the core functionality currently supported.
1. Flexible Resource Management
Data can be kept on-disk to minimize memory usage during processing, or loaded into memory as numpy arrays using .load() for interactive exploration and custom analysis.
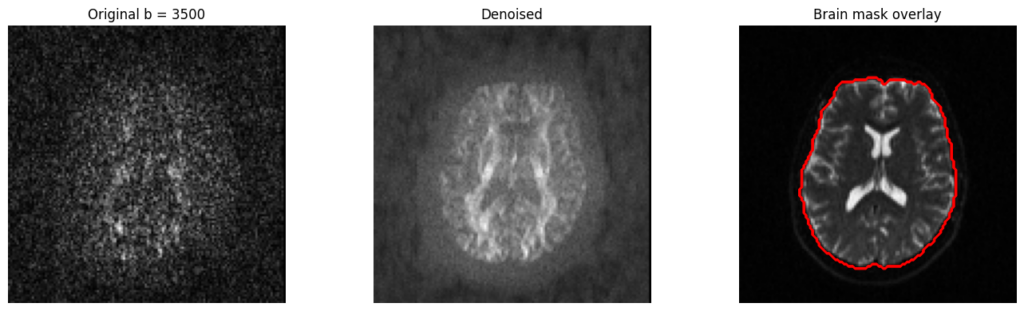
2. Denoising and Brain Extraction
Self-supervised denoising via Patch2Self (DIPY) and deep learning-based brain extraction using HD-BET, with support for both single-subject and batch processing.
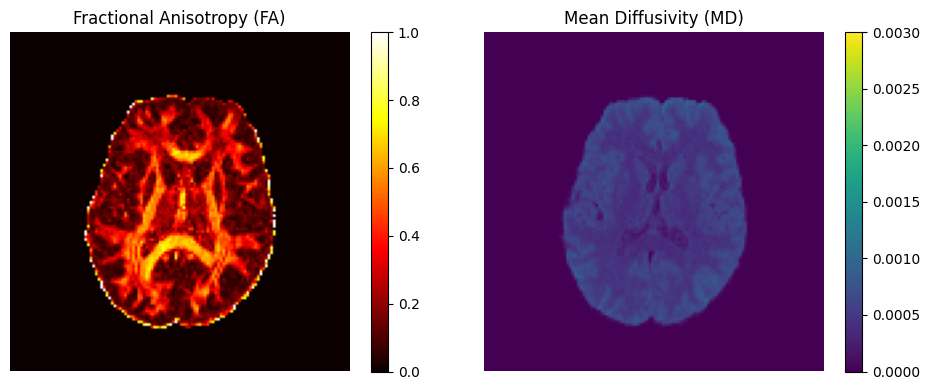
3. Diffusion Tensor Imaging (DTI)
Diffusion tensor modeling using DIPY to compute standard microstructural maps including Fractional Anisotropy (FA) and Mean Diffusivity (MD).
4. Advanced Microstructure modelling
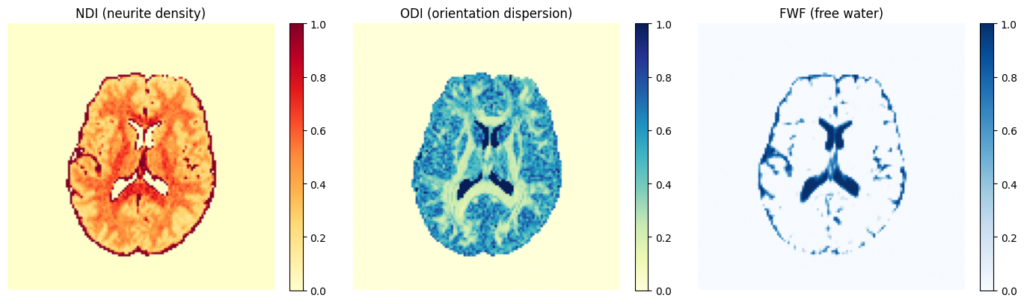
NODDI (Neurite Orientation Dispersion and Density Imaging) estimation using AMICO to extract biophysically-informed metrics of white matter microstructure.
5. Fiber Orientation Modelling
Constrained Spherical Deconvolution (CSD) for estimating fiber orientation distributions (FODs) via DIPY:
- Response function estimation: Our library supports the representation of response functions in spherical harmonic (SH) form with conversion utilities for interoperability between DIPY’s different response formats (prolate tensor and SH-based representations).
- Fiber orientation distribution reconstruction: Computes CSD-based FODs capturing complex fiber configurations within each voxel, and extracts peak directions representing dominant fiber orientations, for use in tractography and connectivity mapping.
- Response function averaging: Population-level averaging following the approach used in MRTrix’s `responsemean` function, which compensates for global magnitude differences between subjects to produce a single group-level response function that serves as a common reference for all FOD estimations.
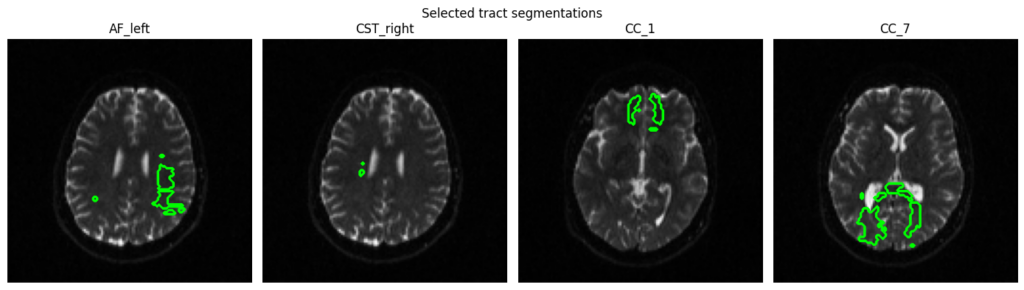
6. Tract Segmentation
Automated white matter tract segmentation via TractSeg using CSD peaks computed with DIPY (rather than MRtrix) to maintain a Python-native implementation.
7. Scalar-Based Registration & Template Building
Registration and template construction via ANTs to facilitate population-level analysis:
- Scalar-based pair-wise registration: Registration between subject pairs driven by any scalar volume, including diffusion metrics (e.g., FA, MD) or standard anatomical T1-weighted images.
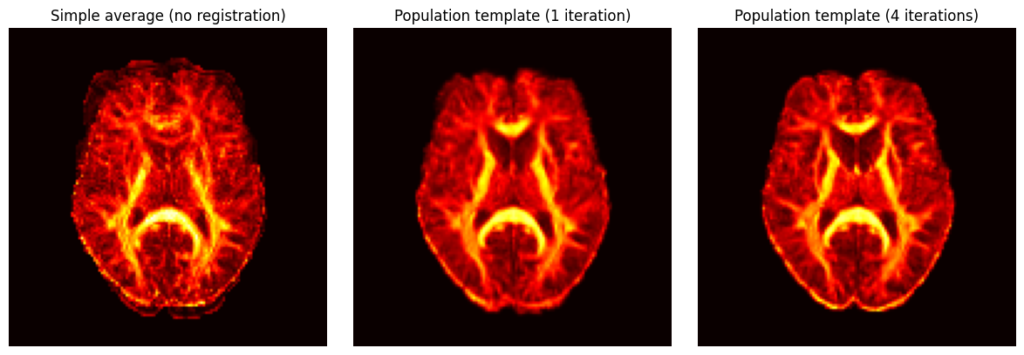
- Single-metric template building: Construction of unbiased population templates following the ANTs iterative template-building approach. Each iteration registers all subjects to the current template using symmetric normalization (SyN), averages the warped images to form an updated template, and applies shape averaging to ensure the template remains unbiased with respect to the input population. Uses a single scalar map per subject (e.g., mean b0, FA or MD).
- Multi-metric template building: Extends the single-metric approach by jointly registering multiple scalar maps from each subject (e.g., mean b0, FA and MD simultaneously) using ANTs multivariate SyN at each iteration. This leverages complementary information from different metrics to achieve more accurate alignment, particularly in regions where individual metrics may provide insufficient contrast.
Try It Out & Collaborate
kwneuro was developed to provide a robust, reproducible and accessible framework for dMRI analysis. Future updates will focus on adding additional pre-processing algorithms (motion correction, gibbs ringing correction) and microstructure models, and adding support for structural MRI. This work is supported by funding for research utilizing the Adolescent Brain Cognitive Development (ABCD) dataset. In future work, we plan to use this library to conduct clinically-relevant analyses of ABCD data to explore developmental brain changes in relation to cognitive and mental health outcomes.
We invite the dMRI community to explore, use, and help shape the future of this library. Our GitHub repository includes interactive Jupyter notebooks demonstrating the library’s functionality, from preprocessing through population-level analysis. Whether you’re a researcher looking to streamline your workflow or a developer interested in contributing novel modeling approaches, we welcome collaboration and feedback. The library is designed to grow with the community’s needs, and we’re committed to making dMRI analysis more accessible and operationally practical for researchers at all levels.
For questions, partnership opportunities, or to learn more about the library’s capabilities, visit https://www.kitware.com/contact or reach out to kitware@kitware.com.
Acknowledgements
This work is supported by the National Institute of Health (NIH) under Award Number 1R21MH132982.