Avogadro and Quantum Chemistry Viz in JupyterLab
We have a new Open Chemistry DOE SBIR Phase II project developing integration of chemical and materials data within Jupyter coupled with a data server/job execution logic. Previously we wrote up some of the early results in a blog post, and presented the platform at JupyterCon in August of this year. In the last few months we have also been looking at making it easier to try some of the tools being developed for this platform. Providing an easy way to install and use some of them in local JupyterLab notebooks is the focus of this post.
Binary Wheels for Avogadro’s Libraries
The Avogadro libraries are lightweight C++ libraries developed to not only support the Avogadro desktop application, but enable reuse in other contexts. We used PyBind11 to wrap some of the core API for reading/writing chemical files, and calculating electronic structure from the output of quantum codes. Binary wheels can be installed using the Python package manager pip under Python 3 (built using our CI integrated with release tags on GitHub):
pip install avogadro
Once you have the package installed you can use a small subset of the C++ API. We plan on extending this in the future, but it is enough to load files, save files, perform some basic electronic structure calculations, and make some very rudimentary queries on the loaded molecule. Our server process has used this for quite a few years, originally using Boost.Python for wrapping.
import avogadro as avo
# Set up the molecule and file format manager
m = avo.core.Molecule()
f = avo.io.FileFormatManager()
# Read an example file
f.read_file(m, 'caffeine.cjson')
# Print some basic attributes
print("Molecule has", m.atom_count(), "atoms,", m.bond_count(),
"bonds, the chemical formula", m.formula(), "and a mass of", m.mass())
# Write a string in the XYZ format
f.write_string(m, "xyz")
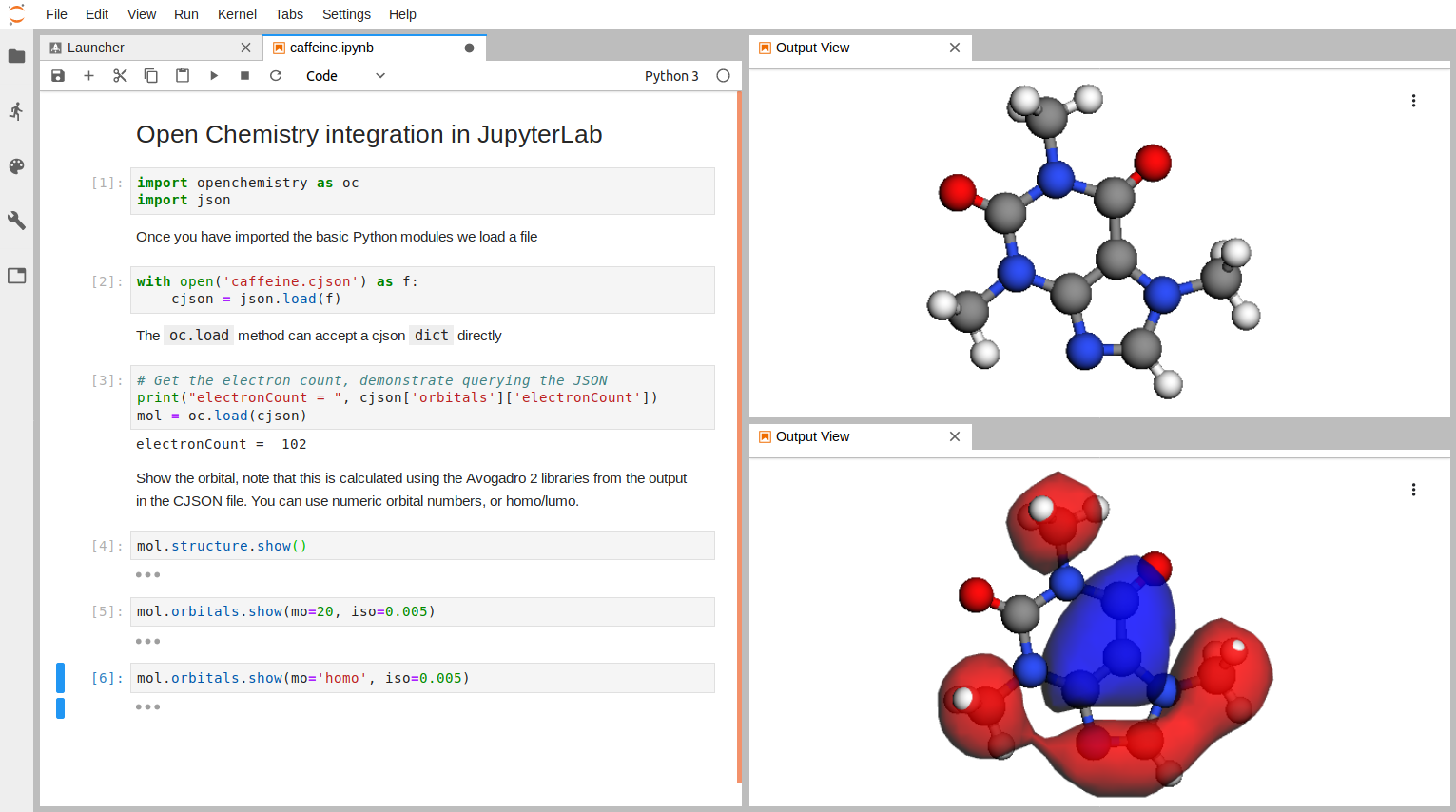
Visualizing Electronic Structure in JupyterLab
Another significant part of our project is the extension of the Jupyter Python 3 kernel, and visualization of chemical data in the JupyterLab web client. We use 3Dmol.js or vtk.js as the JavaScript rendering engines for 3D structure, with D3 for charts. A Python module houses the Python API being developed to support the project, and a JupyterLab extension for the visualization components. These can be installed locally using the following commands assuming you have Python 3 and JupyterLab installed already:
pip install openchemistry jupyter labextension install @openchemistry/jupyterlab
If all goes well you can start your JupyterLab server, and if you clone our new examples repository we will even give you a few example files to get started with:
git clone git://github.com/openchemistry/jupyter-examples jupyter-lab
You can open the example notebooks, and if everything works as expected you will be able to look at the example data, or even import some of your own if you make the files available.
JupyterLab and Binder
We have gone one step further, and added the necessary files to the example repository to enable viewing of these notebooks in Binder. This enables you to experiment with the features described above in a live environment using everything mentioned. It can take a while to launch the containers, but once ready you are able to experiment with no local installation. It is fantastic to be a part of such a vibrant ecosystem of open source tools, and freely available web services that aid in the dissemination of results.
Conclusions
The wrapped Avogadro API is at quite an early stage, and many things you might want to do are not yet supported. Some extra work was done to decouple components so that it is easier to reuse parts of our Python kernel, web visualization, and other pieces independently of the larger project. We hope that you find these pieces useful, and we will continue to work on making it easier to install these open source components independently.
This material is based upon work supported by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences, Small Business Innovation Research (SBIR) program under Award Number DE-SC0017193.